Metagenomics Resource Hub
Featuring workflow validated in real-world hospital
settings and referenced in peer-reviewed clinical studies.
Unified Metagenomic Method for Rapid Detection of Microorganisms in Clinical Samples
We can help. Contact us
We can help. Contact us
Question for the webinar?
Ask here for instant response from this Metagenomics GPT
Ask questions like:
What needs to change in this workflow to apply it to a different sample type?
What was key enabling breakthrough?
What questions were asked at webinar
RECENT PUBLICATIONS
Communications Medicine
Unified Metagenomic Method enabling sameday identification of bacteria, fungi, and both RNA and DNA viruses from asingle clinical sample.
The Lancet Microbe
Unified Metagenomic Method enabling sameday identification of bacteria, fungi, and both RNA and DNA viruses from asingle clinical sample.
Metagenomics Protocol
Complete sample-to-report workflow inunder 7 hours. Pathogen detection after his little as 30 minutes of sequencing
How it works: Sample to report inunder 7 hours.
ArcticZymes powers the upstream host DNA depletion step that makes same-day metagenomics clinically viable.
Watch recent KOL and videodiscussions on clinical metagenomics

From Empiric Guessing to Same-Day Answers: Rethinking Metagenomics for Complicated UTI's [VIDEO]
Read More
Unified Protocol: Rapid Metagenomic Pathogen Detection for ICU Patients [VIDEO]
Read More
How an Accidental Error Helped Unlock Clinical Metagenomics [VIDEO]
Read More
Unlocking Rapid Pathogen Detection with Unified Metagenomics [VIDEO]
Read MoreHost Cell DNA Depletion in Diagnostic Metagenomics
Host nucleic acids often dominateclinical sequencing samples, reducing microbial read depth and limitingpathogen detection. Enzymatic host DNA depletion enriches microbial nucleicacids, improving sequencing sensitivity.
ArcticZymes provides a toolbox ofnucleases designed to efficiently remove host DNA while preserving microbialgenetic material.
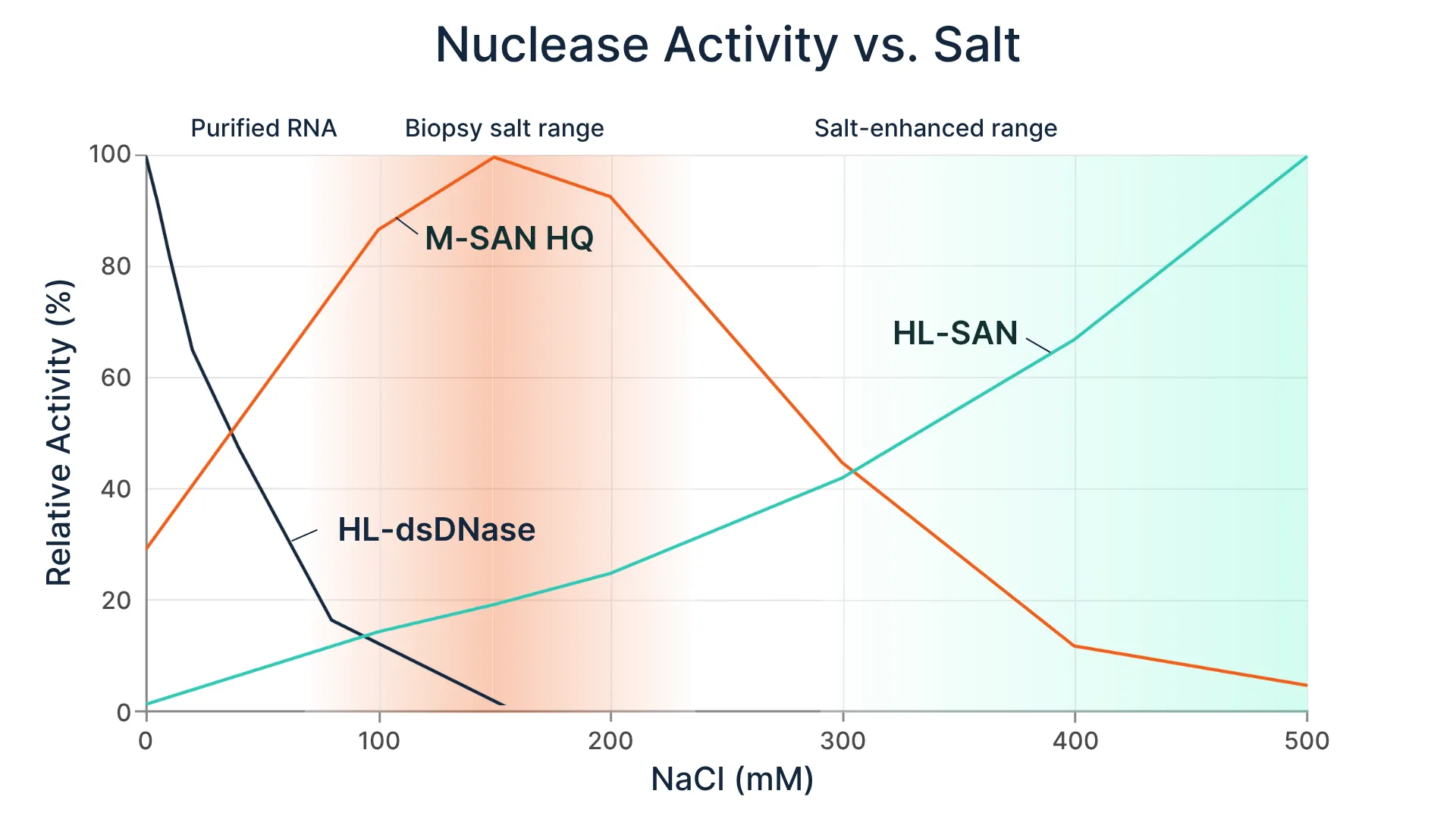
One toolbox. many workflows
Select the appropriate nuclease to tailorhost DNA depletion strategies to your metagenomic workflows.
M-SANHQ
For workflows targeting bothbacterial and viral pathogens, including unified DNA/RNA sequencing approachesand DNA-focused diagnostic applications.
HL-SAN
For workflows that incorporateelevated salt conditions to support effective host DNA removal in moreresilient sample types.
HL-dsDNase
For RNA virus detection workflows where selective removal of contaminating DNA isrequired while preserving RNA.
By selecting the appropriate nuclease from this toolbox, you can tailor host DNA depletion strategies to their metagenomic workflows, improving microbial signal and maximizing sequencing efficiency.

In CSF samples with high level of host DNA, M-SAN HQ ensures efficient host depletion, allowing for enhanced pathogen detection in our diagnostic metagenomic workflow.
- Oslo University Hospital
Join our mailing list for updates
& exclusive metagenomics content


